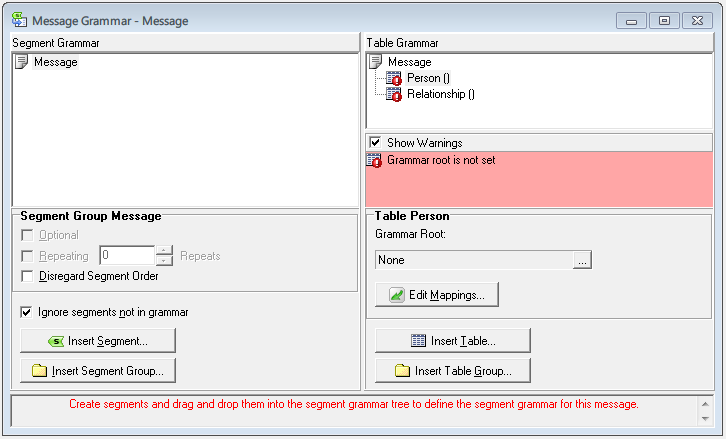
Showing specific residues can realized using: Drawing Method: Isosurface or Volume Slice.Trajectory –> Color Scale Data Range: -5 to +5.Coloring Method: Volume –> with the specified pot….dx.Surface Electrostatics calculated by APBS (stored in a. Graphics Representations… a new window will appear Graphical representation are very versatile in VMD. Save a certain view on your protein Extension Visualization ViewMaster. helix : to select alpha helices exclusivelyĪnd these can be found under the “Singlewords” area.ĭistance Selection: select all FAD that are in a distance of 6 Angstrom from FES select FAD with 6 of resname FESįor aminoacid specific selection Extensions Analysis Sequence Viewer.water : to select for water molecules only.protein : to select for aminoacids only.  
Additional some special selection keyword include chain DĪll these keywords can be found under the “Selection Atoms” tab in the Graphical Representation Windows in the area named “Keywords”. chain : to select for the specific chains e.g.element : to select for the element name e.g.atomicnumber : to select for the atomic number of the atoms e.g.name or type : to select for specific atoms e.g.resid : to select for residues at specific positions e.g.resname : to select for specific residue names (with 3 letter code).Similar to PyMOL, VMD has specific notation to select for certain residues and atoms: Mouse –> Move –> Fragment – 7 – moves fragment by changing its coordinates Mouse –> Move –> Residue – 6 – moves residue by changing its coordinates Mouse –> Move –> Atom – 5 – moves atom by changing its coordinates Mouse –> Label –> Dihedral – 4 – dihedral angle between four atoms Mouse –> Label –> Angle – 3 – angle between three atoms Mouse –> Label –> Bond – 2 – distance between two atoms Query – 0 – information about the picked atom will be displayed in the console Setting the background color to white: color Display Background white To open the console: Extensions Tk Console You would do this by stating explicitly in the Molecule File Browser, Load files for –> select your structure file.Īs in PyMOL all commands in VMD can be realized either by using the graphical interface or the text commands. Here you can either load in another protein file (PDB, PQR, PQRM) or a gridfile, but in the latter you would need to associate it with your structure file. files that indicate position specific properties of your protein or molecule of interest.įiles can be loaded in via the command line: vmd -dx -pdb įiles can also be loaded in via the GUI, using File –> New Molecule… VMD is great for visualizing grid files, i.e. VMD is also special because many other features from other programs in the computational biochemistry field are implemented into VMD. If audio data is included it may be 8-bit raw Pulse-Code Modulation (PCM) samples or 16-bit data encoded with Differential Pulse-Code Modulation (DPCM).VMD stands for visualizing molecular dynamics and is besides PyMOL one of the most used molecular visualization softwares out there, at least for proteins. Video data is encoded with the Lempel-Ziv (LZ77) algorithm and run-length encoding (RLE). The VMD format begins with a header that includes information about the video, such as the video codec version, the width and height of the video frame, and the audio sample rate. Many of the video games that utilize this format are interactive games that feature a large number of cutscenes with human actors in real or 3D animated environments. VMD was developed by Sierra On-Line (now Sierra Entertainment) in the 1990s to store video featured in their CD-ROM video games. Some games that use VMD files besides Phantasmagoria include Betrayal in Antara, SWAT, RAMA, Shivers, Lighthouse, The Last Dynasty, and Torin's Passage. You will most likely only encounter a VMD file if you have installed a game that utilizes the TGV format on your computer. VMD file open in VideoLAN VLC media player 3
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
